Working with data¶
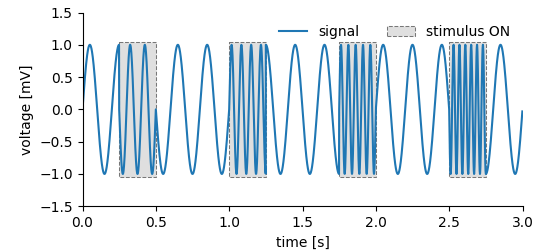
Storing data is one thing, but we want to work with it. The following examples illustrate the reading of data from DataArray, Tag and MultiTag entities We will use the dummy dataset already used in the tagging example.
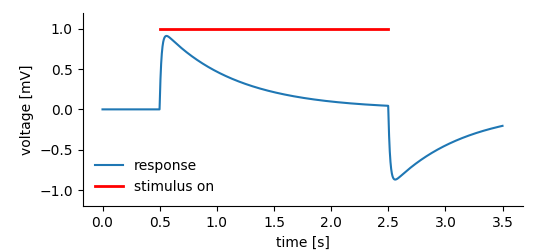
The following code creates the dummy data and links the Tag providing the context (a stimulus was on in a certain time segment) to the DataArray that actually stores the data.
#include <nix.hpp>
#include <numeric>
int main() {
double interval = 0.01;
double stim_on = 0.5;
double stim_off = 2.5;
std::vector<double> response(350);
std::iota(response.begin(), response.end(), 0.);
std::transform(response.begin(), response.end(), response.begin(),
[interval](double x){ return x * interval; });
std::transform(response.begin(), response.end(), response.begin(),
[stim_on, stim_off](double x){ return (x >= stim_on && x < stim_off) ? 1 : 0; });
nix::File f = nix::File::open("tagging1.nix", nix::FileMode::Overwrite);
nix::Block block = f.createBlock("demo block", "nix.demo");
nix::DataArray data = block.createDataArray("response", "nix.sampled", response);
data.label("voltage");
data.unit("mV");
nix::SampledDimension dim = data.appendSampledDimension(interval);
dim.label("time");
dim.unit("s");
nix::Tag stim_tag = block.createTag("stimulus", "nix.stimulus_segment", {stim_on});
stim_tag.extent({stim_off - stim_on});
stim_tag.addReference(data);
f.close();
return 0;
}
In this example we know the interesting entities by name, i.e. the DataArray is called response and the Tag is called stimulus. In cases in which we have no clue about the names, we just have to browse the file or search by name or type.
Reading data¶
The first and maybe most common problem is to read the data stored in a DataArray.
Reading all data¶
In NIX when you open a DataArray the stored the data is not automatically read from file. This needs to be done separately by performing the following steps.
Open the file, get the Block and the DataArray
Get the data from file.
Get the timestamps from the dimension.
#include <nix.hpp>
#include <boost/optional/optional_io.hpp> // only needed for << operator on optionals
void main() {
// 1. Open file, block, and dataarray
nix::File file = nix::File::open("tagging1.nix", nix::FileMode::ReadOnly);
nix::Block block = file.getBlock("demo block");
if (!block.hasDataArray("response"))
return -1;
nix::DataArray responseArray = block.getDataArray("response");
// 2. read the data to the vector
std::vector<double> responseData;
responseArray.getData(responseData);
// 3. get the timestamps
SampledDimension dim = responseArray.getDimension(1).asSampledDimension();
std::vector<double> time = dim.axis(responseData.size());
// some output to confirm it worked
std::cerr << dim.label() << "\t" << responseArray.label() << std::endl;
std::cerr << dim.unit() << "\t" << responseArray.unit() << std::endl;
for (size_t i = 0; i < responseData.size(); ++i) {
std::cerr << time[i] << "\t" << responseData[i] << std::endl;
}
file.close();
return 0;
}
There are a few noteworthy things:
We use some previous knowledge here. For one, we know the names of the entities. Next, we know the data type of the data (double). Further, we know that the data is 1-D and the single dimension is a
SampledDimension. If these things are not known, the NIX library offers the necessary functions to get this information.DataArray::dataExtent()returns annix::NDSizethat is the data extent/shape.DataArray::dataType()returns the data type.To find out the
DimensionType, we need to do something like:
nix::Dimension dim = responseArray.getDimension(1); if (dim.dimensionType() == nix::DimensionType::Sampled) nix::SampledDimension sd = dim.asSampledDimension();
Note: Dimension indices start with 1.
We use a std::vector, this is not necessary. We could also use boost mulitarrays, for example.
DataArrat::getData()will automatically resize the vector. When reading all the data we do not need to worry about the data extent.DataArray and Dimension functions
label()andunit()returnboost::optionalvalues. Since these fields are optional, the returned optionals may not contain values. Here, we use them only for output (and thus need to include the boost header).
Reading partial data¶
In other instances it might be wanted to read only parts of the data.
When doing this, the library needs to know where it should start to read
and how many elements should be read. Because DataArrays can store
n-dimensional data we need to specify these with an n-dimensional
construct, the nix::NDSize.
The following steps need to be taken.
Open the file, get the Block and the DataArray
Define offset and count.
Read the data from file.
Get the timestamps from the dimension.
#include <nix.hpp>
#include <boost/optional/optional_io.hpp> // only needed for << operator on optionals
void main() {
// 1. Open file, block, and dataarray
nix::File file = nix::File::open("tagging1.nix", nix::FileMode::ReadOnly);
nix::Block block = file.getBlock("demo block");
if (!block.hasDataArray("response"))
return;
nix::DataArray responseArray = block.getDataArray("response");
// 2. define offset and count
nix::NDSize dataExtent = responseArray.dataExtent();
nix::NDSize offset(1, (int)(dataExtent[0]/4));
nix::NDSize count(1, (int)(dataExtent[0]/2));
// 3. read the data
std::vector<double> responseData;
responseArray.getData(responseData, count, offset);
// 4. get the timestamps
SampledDimension dim = responseArray.getDimension(1).asSampledDimension();
std::vector<double> time = dim.axis(count[0], offset[0]);
// some output to confirm it worked
std::cerr << dim.label() << "\t" << responseArray.label() << std::endl;
std::cerr << dim.unit() << "\t" << responseArray.unit() << std::endl;
for (size_t i = 0; i < responseData.size(); ++i) {
std::cerr << time[i] << "\t" << responseData[i] << std::endl;
}
file.close();
return 0;
}
Again, some things may be noted here: 1. We use some previous knowledge
in this example. 2. We are reading half of the data and start at 1/4th
of the data extent. 3. We read to a std::vector that will be resized
appropriately. 4. We use the axis() function of the
SampledDimension to get matching timestamps that match offset and
count.